The \(UF_6\) anion
The problem
This case study, suggested by Kirk Peterson concerns the calculation of the \(UF_6\) anion using effective core potentials (ECP)
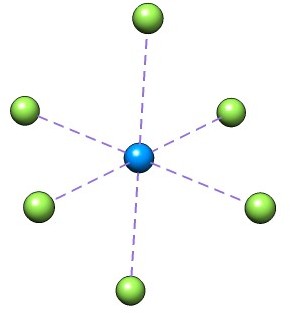
We start from the molecular input UF6.mol
INTGRL
UF6- vqz geom
cc-pVDZ-PP
C 2
92. 1
U 0.0 0.0 0.0
LARGE BASIS cc-pVDZ-PP
ECPLIB ECP60MDF
9. 6
F 0.0000000 0.0000000 3.9259448
F 3.9259448 0.0000000 0.0000000
F 0.0000000 -3.9259448 0.0000000
F -3.9259448 0.0000000 0.0000000
F 0.0000000 3.9259448 0.0000000
F 0.0000000 0.0000000 -3.9259448
LARGE BASIS aug-cc-pVDZ
FINISH
and the menu file UF6.inp
**DIRAC
.TITLE
UF6-
.WAVE FUNCTION
.ANALYZE
**ANALYZE
.MULPOP
**HAMILTONIAN
.ECP
**INTEGRALS
*READIN
.UNCONTRACT
**WAVE FUNCTION
.SCF
*SCF
.CLOSED SHELL
44 42
.OPEN SHELL
1
1/0,14
.MAXITR
75
*END OF
The problem is that this calculation simply does not converge.
Analysis
The neutral molecule is a simple closed shell molecule and using the menu file
**DIRAC
.TITLE
UF6-
.WAVE FUNCTION
.ANALYZE
**HAMILTONIAN
.ECP
**GENERAL
.ACMOUT
**ANALYZE
.MULPOP
*MULPOP
.VECPOP
1..40
1..40
**INTEGRALS
*READIN
.UNCONTRACT
**WAVE FUNCTION
.SCF
*SCF
.CLOSED SHELL
44 42
.MAXITR
75
*END OF
the SCF converges nicely in 26 iterations. Before proceeding we investigate the electronic structure of this species using projection analysis, see [Dubillard2006] and this tutorial. Looking at the input file above, one may note that we already prepared for this by invoking the keyword .ACMOUT in order to recover the \(C_1\) MO coefficients:
pam --inp=UF6 --mol=UF6 --outcmo --get "DFACMO=ac.UF6"
We now generate orbitals for the constituent atoms of \(UF_6\) in their electronic ground state.
For the fluorine atoms this is straightforwardly obtained using the molecular input F.mol
INTGRL
UF6- vqz geom
cc-pVDZ-PP
C 1
9. 1
F 0.0000000 0.0000000 0.0000000
LARGE BASIS aug-cc-pVDZ
FINISH
and the menu file F.inp
**DIRAC
.WAVE FUNCTION
.ANALYZE
**GENERAL
.ACMOUT
**HAMILTONIAN
.X2C
**INTEGRALS
*READIN
.UNCONTRACT
**WAVE FUNCTION
.SCF
*SCF
.CLOSED SHELL
4 0
.OPEN SHELL
1
5/0,6
**END OF
using the command:
pam --inp=F --mol=F --outcmo --get "DFACMO=ac.F"
mv DFCOEF cf.UF6
Note that since no effective core potential has been given for the fluorine we prefer to calculate it using the X2C Hamiltonian.
The corresponding molecular input for the uranium atom is
INTGRL
UF6- vqz geom
cc-pVDZ-PP
C 1
92. 1
U 0.0 0.0 0.0
LARGE BASIS cc-pVDZ-PP
ECPLIB ECP60MDF
FINISH
For the uranium atom, the calculation of the occupation easily converges by using the keyword .KPSELE introduced in DIRAC21.
**DIRAC
.WAVE FUNCTION
.ANALYZE
**GENERAL
.ACMOUT
**HAMILTONIAN
.ECP
**INTEGRALS
*READIN
.UNCONTRACT
**WAVE FUNCTION
.SCF
*SCF
.CLOSED SHELL
16 12
.OPEN SHELL
2
3/0,14
1/10,0
.KPSELE
7
-1 1 -2 2 -3 3 -4
4 4 8 4 6 0 0
0 0 0 0 0 6 8
0 0 0 4 6 0 0
**ANALYZE
.MULPOP
*MULPOP
.VECPOP
1..oo
1..oo
**END OF
using the command:
pam --inp=U --mol=U --get "DFACMO=ac.U"
The projection analysis is carried out in \(C_1\) symmetry to make all fluorines symmetry-independent. The proper molecular input for \(UF_6\) is found as an xyz-file in the DIRAC output
7
U 0.0000000000 0.0000000000 0.0000000000
F 0.0000000000 0.0000000000 2.0775205092
F 0.0000000000 0.0000000000 -2.0775205092
F 2.0775205092 0.0000000000 0.0000000000
F -2.0775205092 0.0000000000 0.0000000000
F 0.0000000000 2.0775205092 0.0000000000
F 0.0000000000 -2.0775205092 0.0000000000
whereas the menu file prj.inp is given by
**DIRAC
.TITLE
UF6-
.ANALYZE
**HAMILTONIAN
.ECP
**ANALYZE
.PROJEC
*PROJEC
.OWNBAS
.VECPRJ
1..50
.VECREF
7
AFUXXX
1
1..26
AFF1XX
1
1..5
AFF2XX
1
1..5
AFF3XX
1
1..5
AFF4XX
1
1..5
AFF5XX
1
1..5
AFF6XX
1
1..5
**INTEGRALS
*READIN
.UNCONTRACT
**WAVE FUNCTION
.SCF
*SCF
.CLOSED SHELL
86
.MAXITR
75
**MOLECULE
*SYMMETRY
.NOSYM
*BASIS
.SPECIAL
F BASIS aug-cc-pVDZ
.DEFAULT
cc-pVDZ-PP
ECPLIB ECP60MDF
*END OF
Before running the analysis we have to create symbolic links for the reference files:
ln -s ac.U AFUXXX
ln -s ac.F AFF1XX
ln -s ac.F AFF2XX
ln -s ac.F AFF3XX
ln -s ac.F AFF4XX
ln -s ac.F AFF5XX
ln -s ac.F AFF6XX
The analysis is now run using:
cp ac.UF6 DFCOEF
pam --inp=prj --mol=UF6.xyz --incmo --copy="AF*"
Looking at the end of the output we find
* Total gross contributions:
AFUXXX 28.4581 E - 28.4581 P - 0.0000
AFF1XX 9.5325 E - 9.5325 P - 0.0000
AFF2XX 9.5325 E - 9.5325 P - 0.0000
AFF3XX 9.5325 E - 9.5325 P - 0.0000
AFF4XX 9.5325 E - 9.5325 P - 0.0000
AFF5XX 9.5325 E - 9.5325 P - 0.0000
AFF6XX 9.5325 E - 9.5325 P - 0.0000
Polarization: 0.3472
showing that the polarization contribution is rather small. It can be eliminated completely
by adding the keyword .POLREF under the *PROJEC section (valid from DIRAC13).
The gross population of the uranium valence orbitals is then given by
* Total reference orbital contributions:
AFUXXX E 1 2.00002 -0.12565689E+02
AFUXXX E 2 2.00000 -0.10085084E+02
AFUXXX E 3 1.99999 -0.80747936E+01
AFUXXX E 4 1.99999 -0.80747936E+01
AFUXXX E 5 1.99934 -0.43446774E+01
AFUXXX E 6 1.99934 -0.43446774E+01
AFUXXX E 7 1.99905 -0.40406678E+01
AFUXXX E 8 1.99948 -0.40406678E+01
AFUXXX E 9 1.99913 -0.40406678E+01
AFUXXX E 10 1.99055 -0.21324652E+01
AFUXXX E 11 1.99674 -0.13354477E+01
AFUXXX E 12 1.97251 -0.98353353E+00
AFUXXX E 13 1.97251 -0.98353353E+00
AFUXXX E 14 0.24253 -0.34920842E+00
AFUXXX E 15 0.11154 -0.34920842E+00
AFUXXX E 16 0.26380 -0.34920842E+00
AFUXXX E 17 0.19815 -0.32405790E+00
AFUXXX E 18 0.09020 -0.32405790E+00
AFUXXX E 19 0.12766 -0.32405790E+00
AFUXXX E 20 0.20009 -0.32405790E+00
AFUXXX E 21 0.06505 -0.20167848E+00
AFUXXX E 22 0.28390 -0.19291378E+00
AFUXXX E 23 0.28390 -0.19291378E+00
AFUXXX E 24 0.25493 -0.18402313E+00
AFUXXX E 25 0.24577 -0.18402312E+00
AFUXXX E 26 0.25560 -0.18402312E+00
...
* Total gross contributions:
AFUXXX 28.5518 E - 28.5518 P - 0.0000
AFF1XX 9.5747 E - 9.5747 P - 0.0000
AFF2XX 9.5747 E - 9.5747 P - 0.0000
AFF3XX 9.5747 E - 9.5747 P - 0.0000
AFF4XX 9.5747 E - 9.5747 P - 0.0000
AFF5XX 9.5747 E - 9.5747 P - 0.0000
AFF6XX 9.5747 E - 9.5747 P - 0.0000
Polarization: 0.0000
We can identify the various orbitals from orbital energies and degeneracies or by looking in the Mulliken output of the atomic calculation. It is seen that the fluorines have charges -0.57 and uranium +3.45. The electronic configuration of uranium in the neutral molecule is \(5f^{1.23}6d^{1.32}7s^{0.06}\).
Calculating the anion
Now let us consider adding an electron. From the output of the neutral molecule we find that the HOMO
(ungerade) has energy -0.642 \(E_h\) and the LUMO (ungerade) comes in at -0.115 \(E_h\).
In fact, Mulliken population analysis shows that the first seven virtual orbitals are indeed basically f orbitals.
We therefore attempt a straightforward calculation starting from the coefficients of the neutral molecule
and using the menu file UF6a.inp
**DIRAC
.TITLE
UF6-
.WAVE FUNCTION
.ANALYZE
**HAMILTONIAN
.ECP
**ANALYZE
.MULPOP
*MULPOP
.VECPOP
1..oo
1..oo
.LABEL
SHELL
**INTEGRALS
*READIN
.UNCONTRACT
**WAVE FUNCTION
.SCF
*SCF
.CLOSED SHELL
44 42
.OPEN SHELL
1
1/0,14
.OVLSEL
.NODYNSEL
*END OF
and the command:
pam --copy="cf.UF6=DFCOEF" --inp=UF6a --mol=UF6 --outcmo
Note that we activate static overlap selection to keep occupied orbitals in place.
However, this calculation does not converge. Also, in the Mulliken population analysis
we activate the SHELL keyword to simplify identification of atomic orbitals, see
.LABEL
Looking at the open-shell orbitals we find that they are essentially f orbitals. However, we also note that the HOMO of the closed shell orbitals comes in at -0.40353915 \(E_h\) and the lowest occupied open-shell orbitals is almost degenerate, having orbital energy -0.39634956 \(E_h\). Clearly this almost degeneracy creates artificial mixing which perturbs convergence. We therefore add an open-shell level shift:
.OLEVEL
0.200
and now the SCF converges in 38 iterations. Curiously, convergence is extremely sensitive to the choice of the level shift. Even changing it by 0.005 breaks convergence !
- orphan:
The \(UF_6\) anion - another approach
Different and more robust solution for obtaining the SCF convergence of the \(UF_6\) anion is provided here. It is based on the damping of the Fock matrix from SCF iteration to iteration.
Starting from neutral \(UF_6\)
In the first step we follow the previous approach of starting from the closed-shell (neutral) \(UF_6\) system.
(We also may utilize a shift in the virtual space to reduce mixing between occupied and virtual spinors.) In any case, we
get smooth convergence with the default DIIS method. Files to download are UF6neutral.inp, UF6.mol,
cc-pVDZ-PP and ECP60MDF:
pam --noarch --inp=UF6neutral.inp --mol=UF6.mol --get "DFCOEF=DFCOEF.UF6neutral"
The \(UF_6\) anion from neutral
In the next step we compare two calculations starting from neutral molecular spinors of UF6 (the file DFCOEF.UF6neutral).
One calculation uses the DIIS method (not shown here) and the other uses the damping with two damping factors of 0.90 and 0.50.
(File to download is UF6_anion_scfdamp.inp):
pam --noarch --inp=UF6_anion_scfdamp.inp --mol=UF6.mol --put "DFCOEF.UF6neutral=DFCOEF" --get "DFCOEF=DFCOEF.UF6anion_damp_pass1" --replace dampfactor=0.9
pam --noarch --inp=UF6_anion_scfdamp.inp --mol=UF6.mol --put "DFCOEF.UF6anion_damp_pass1=DFCOEF" --get "DFCOEF=DFCOEF.UF6anion_damp_pass2" --replace dampfactor=0.5
Concerning the DIIS speedup method, the SCF iterations initially converge to ca \(10^{-5}\) but then began to diverge and after about 150 iterations the error gradient of the Fock matrix is still around \(10^{-5}\).
However, with the damping convergence speedup the calculation converges to ca \(10^{-8}\) after about 50 iterations. The damping scheme uses more of the input Fock matrix that the output Fock matrix. Concerning the damping factor, the lower than 1 it is, we should get faster convergence. However, when the factor is getting too small, iterations diverge. The closer we are to the converged states, the smaller the damping factor can be.
If we begin calculations from the converged solutions from the damping, the DIIS iterations converge
rapidly after few iterations. But this is only with the deactivated active-active shells interaction
(new file to download is UF6anion2.inp):
pam --noarch --inp=UF6anion2.inp --mol=UF6.mol --put "DFCOEF.UF6anion_damp_pass2=DFCOEF" --get "DFCOEF=DFCOEF.UF6anion_diis"
The \(UF_6\) anion with OPENFAC=1
Unfortunately, with the factor OPENFAC=1 (what is the keyword .OPENFACTOR) we were not able to get the convergence,
neither with the default DIIS, nor with damping factor (to download, UF6anion.inp and UF6anion3.inp):
pam --noarch --inp=UF6anion.inp --mol=UF6.mol --put "DFCOEF.UF6anion_diis=DFCOEF"
pam --noarch --inp=UF6anion3.inp --mol=UF6.mol --put "DFCOEF.UF6anion_diis=DFCOEF" --replace dampfactor=0.8
More effort is required to obtain converged molecular spinors of the \(UF_6\) anion with the active-active open-shell interaction (that is OPENFAC=1). We leave this task to the reader.